Introduction
Hepatocellular carcinoma (HCC), a highly invasive and heterogeneous malignancy, is the third leading cause of cancer deaths globally (~830,000 annually), with 50% of cases linked to HBV infection [1–3]. Despite treatments like surgery, transplantation, and targeted therapy, the 5-year survival rate remains ≤ 18% due to tumour heterogeneity and drug resistance [4–6]. Traditional experimental models for HCC research (e.g. cell lines, patient-derived xenografts [PDX]) fail to fully replicate the tumour microenvironment (TME) or meet research needs efficiently [7, 8].
Organoid technology has emerged as a transformative tool, enabling 3D culture of stem cells or patient-derived tissues to mimic TME, retain tumour biology, and predict drug responses (accuracy up to 80%) [9, 10]. However, rapid growth in this field has led to fragmented knowledge, uneven resource allocation, and limited interdisciplinary collaboration, hindering critical breakthroughs [11, 12].
To address this, we conducted a bibliometric analysis of HCC organoid research (2005–2024) using the Web of Science Core Collection (WOSCC) data, VOSviewer, and CiteSpace [13, 14]. The study aimed to map the field’s evolution, to identify frontiers, and to provide insights for new researchers, aiming to guide future advancements [15].
Aim
This bibliometric study examines HCC organoid research evolution and emerging trends over two decades.
Material and methods
Data sources and search strategy
The data used in this study are derived from the WOSCC database. The search strategy is presented in Supplementary Table SI.
Literature screening and data extraction
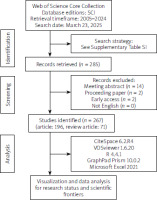
This study used the WOSCC as the only data source. After systematic screening from 1 January 2005 to 31 December 2024, we removed meeting abstracts (14 articles), early access literature (2 articles), and proceeding papers (2 articles) from the initial 285 articles, retaining 267 articles (196 articles and 71 review articles). All literatures were uniformly preserved in a ‘Plain Text File’ format on 23 March 2025. The screening process is shown in Figure 1.
Data analysis
Bibliometric analysis was performed using CiteSpace (6.2.R4) for keyword burst, journal double map overlay, cooperative network analysis, and VOSviewer (1.6.20) constructing a co-occurrence network. The ‘ggplot2’, ‘pheatmap’, and ‘circlize’ packages of R language (4.4.1) were used to draw a geographic distribution map, heat map, and chord diagram, respectively. Data were analysed using Microsoft Excel 2021 and GraphPad Prism (10.0.2).
Results
Annual publication trends
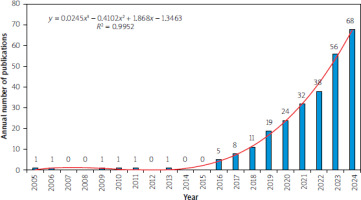
The HCC organoid model showed significant stage development characteristics (Figure 2). From 2005 to 2015, the average annual number of published papers was less than 1. Since 2016, it entered a period of rapid growth, exceeding 10 papers in 2018 (11 papers). It is worth noting that the number of publications in 2018–2024 accounted for 92.88% of the total.
Country/region contribution analysis
Global HCC organoid research involved 40 countries/regions (Figure 3 A). The data showed that China ranked first, with 128 articles, followed by the United States (65 articles), Germany (27 articles), and the Netherlands (24 articles) (Table I and Supplementary Table SII). Although England ranked seventh in the number of publications (10 articles), its average citation rate reached 144.8, the highest among all the countries. Japan (19 articles) and South Korea (8 articles) in East Asia had high research activity, Australia and Egypt each contributed 6 articles, Russia only 1 article, and African countries had relatively low participation.
Figure 3
A – Global geographical distribution map. B – Top 15 countries/regions annual publication line chart. C – Annual heat map of the top 15 countries/regions D – The country/region cooperation network diagram based on CiteSpace, which mainly highlights the cooperation relationship. E – Based on VOSviewer’s country/region cooperation network diagram, it mainly emphasises the intensity of cooperation. The node represents the country/region, the size of the node reflects the number of publications, and the thickness of the connection indicates the intensity of cooperation

Table I
Top 15 countries/regions by research contribution to HCC organoids based on publication output, citation impact, and average citation
The annual trend analysis of Figures 3 B and C shows that the research output of China and the United States in this field shows a continuous growth trend. The cooperation network diagram (Figures 3 D, E) constructed by CiteSpace and VOSviewer shows that China and the United States are at the centre of the global cooperation network, and the cooperation intensity between the two countries is the highest. European countries such as Germany and the Netherlands, maintain close scientific research cooperation with China and the United States.
Institutional contribution analysis
CiteSpace analysed 630 HCC organoid research institutions. As Figure 4 A and Table II show, Chinese institutions dominate this field. Naval Medical University (18 papers, 26.94%), Chinese Academy of Sciences (15 papers, 29.53%), and Sun Yat Sen University (14 papers, 38.21%) ranked the top three in terms of the number of papers published, and Chinese institutions accounted for six of the top ten. It is particularly noteworthy that although Shanghai Jiao Tong University has a small number of publications, its annual average citation rate of 64.11% highlights its outstanding academic influence.
Figure 4
A – Institutional cooperation network diagram. B – The cooperation strength chord diagram of the top 10 contributing institutions C – The author cooperation network diagram. D – Author co-citation network diagram

Table II
Top 10 institutions contributing to HCC organoid research: publication output and citation impact
Further analysis of the cooperation intensity of the top ten institutions through the ‘circlize’ package of R language (version 4.4.1) (Figure 4 B) shows that Naval Medical University, Chinese Academy of Sciences, and Fudan University have formed a close cooperation network. In contrast, the Chinese Academy of Medical Sciences – Peking Union Medical College, Ruprecht Karls University Heidelberg in Germany, and the University of California system have not yet established significant cooperative relationships with other institutions.
Author and co-cited author analysis
This study analysed the contributions of 2315 authors in the field of HCC organoids. Table III shows that 61 papers (22.85% of the total) were published by the top ten authors, including nine by Van Der Laan, Luc J. W. and Verstegen, Monique M. A., and six by Ma, Stephanie.
Table III
Top 10 authors and co-cited authors of HCC organoid studies: number of publications and citations
Through the analysis of the cooperation network of the top 50 high-yield authors using VOSviewer (Figure 4 C), the researchers formed eight distinguished cooperative clusters. This relatively independent clustering model shows that research in this field has been divided into multiple characteristic directions, and the cooperation within each team is close; however, the cross-team cooperation is limited.
The co-citation analysis (Figure 4 D and Table III) showed that Broutier L (108 citations), Llovet Jm (78 citations), and Huch M (66 citations) were the most cited authors in this field, constituting the core knowledge base of the research.
Analysis of academic journals
This study systematically analysed the publications of journals in the field of HCC organoids. According to the statistics of 149 journals, the top ten are all JCR Q1 partitions, which shows that the overall research level is high (Table IV). International Journal of Molecular Sciences ranked first with 11 articles (4.12%), followed by Cancer and Hepatology with 9 articles (3.37%). Although the number of articles published in the Journal of Hepatology is not among the top three, its 26.8 impact factor highlights its outstanding academic influence.
Table IV
Top 10 journals in the field of HCC organoid research published papers.
The analysis of the journal cooperation network (Figure 5 A) shows that although journals such as Nature Medicine and Cell Reports have not published a large number of papers, they are at the centre of the network, showing the ability of cross-journal knowledge exchange and playing an important role as academic bridges.
Figure 5
A – Journal cooperation network diagram A B – Journal co-citation network diagram. C – Journal double overlay map (Left: citing journals. Right: Cited journals)

The co-citation analysis of journals (Table V and Figure 5 B) showed that Cell ranked first with 191 citations, followed by Hepatology (182) and Nature (178), which together constitute the theoretical basis of the field. Although the citation frequency of Natural Medicine is not high, the impact factor of 58.7 still reflects an extremely high value.
Table V
Top 10 co-cited journals in HCC organoid research
The double overlay diagram of journals (Figure 5 C) shows that the cited journals are mostly from the fields of molecular, biology, and genetics, whereas the citing journals are concentrated in the fields of molecular, biology, and immunology. The analysis of cross-citation patterns between the two journal clusters reveals limited interdisciplinary integration in this research field.
Key word and trend topic analysis
We used VOSviewer to construct a co-occurrence network of 183 keywords (minimum frequency = 3; Figure 6 A). Supplementary Table SIII lists the top 20 high-frequency keywords, including ‘hepatocellular carcinoma’, ‘organoid’, ‘expression’, ‘cancer’, ‘sorafenib’, ‘in vitro’, ‘stem cells’, ‘disease’, ‘activation’, and ‘cells’, reflecting the core concepts in this field. The keyword network overlap map and volcano map (Figures 6 B and C) show the research hotspots and their evolution trends, such as the prominent position of keywords such as ‘b-catenin’, ‘liver cancer’, ‘DNAJB1-PRKACA’, ‘animal models’, ‘double blind’, ‘drug resistance’, ‘1st-line treatment’, and ‘targeted therapy’.
Figure 6
A – Keyword co-occurrence network. B – Network overlap of co-occurrence keywords C – Keyword volcano map. Volcanoes correspond to hot topics in each time period. D – The diagram of top 30 keywords with the strongest citations bursts

Key word burst analysis provides valuable insights and academic guidance for researchers to track the dynamic evolution of hotspots. Figure 6 D shows 30 keywords with the strongest citation bursts. Established concepts like ‘TGF-b’ and ‘apoptosis’ have maintained sustained scholarly attention. Notably, emerging terms including ‘long-term expansion’, ‘fibrosis’, ‘patient-derived organoids’, ‘oxidative stress’, ‘primary liver cancer’, ‘b-catenin’, and ‘acid’ exhibit recent surge patterns, signalling growing research interest that may define future investigative priorities in HCC organoid research.
Analysis of co-cited reference
Figure 7 A shows the co-cited reference network map of HCC organoid research from 2005 to 2024, which contained 535 nodes and 2868 links. Supplementary Table SIV lists the 10 key studies with the highest citation frequency, among which ‘Human primary liver cancer-derived organoid cultures for disease modelling and drug screening’ published by Nature Medicine ranked first, with 52 citations. ‘Organoid Models of Human Liver Cancers Derived from Tumour Needle Biopsies’ published in Cell Reports was followed by 42 citations. ‘Human primary liver cancer organoids reveal intratumor and interpatient drug response heterogeneity’ published by JCI Insight ranked third with 38 citations.
Figure 7
A – Co-cited reference network map. B – Keyword clustering diagram of co-cited reference. C – Co-cited reference volcano map. D – Top 40 references with the strongest citation bursts

In addition, co-cited reference keyword cluster analysis and volcano mapping were performed using CiteSpace (Figures 7 B and C). Early research hotspots were identified as leucine-rich repeat-containing G-protein-coupled receptor 5 (LGR5) (cluster 2) and epithelial-to-mesenchymal transition (cluster 4). The mid-term research focus was dominated by single-cell-based analysis (cluster 3), liver models (cluster 7), animal models (cluster 13), and biliary tree stem cells (cluster 14).
Figure 7 D shows the top 40 articles with the highest number of citation bursts. The top three most cited reports include Broutier et al.’s ‘Human primary liver cancer-derived organoid cultures for disease modeling and drug screening’ (strength = 13.7513), Huch et al.’s ‘Long-Term Culture of Genome-Stable Bipotent Stem Cells from Adult Human Liver’ (strength = 7.4748), and Van De Wetering et al.’s ‘Prospective Derivation of a Living Organoid Biobank of Colorectal Cancer Patients’ (strength = 6.0585).
Discussion
Visual analysis of keywords in this study illustrates the current hotspots and emerging trends in the field of HCC organoids. The network of 183 co-occurrence keywords highlighted terms such as ‘hepatocellular carcinoma’, ‘organoid’, ‘expression’, ‘cancer’, ‘sorafenib’, ‘in vitro’, ‘stem-cells’, ‘disease’, ‘activation’, and ‘cells’. At the same time, the analysis of keyword timeline highlights terms such as ‘b-catenin’, ‘liver cancer’, ‘the DNAJB1-PRKACA fusion- the signature genetic event of fibrolamellar hepatocellular carcinoma (FL-HCC)’, ‘animal models’, ‘double blind’, ‘drug resistance’, ‘1st-line treatment,’ and ‘targeted therapy’.
In terms of research hotspots, the most prominent direction is the construction and optimisation of HCC organoid models. Several studies have shown that researchers have a strong interest in the establishment of organoid culture systems of different sources and types, such as patient tissues, stem cells, primary cancer, fibrolamellar carcinoma, and metabolic-related cancer [16–21]. In particular, the modelling of rare subtypes of fibrolamellar carcinoma provides a valuable platform for drug testing and development [22].
Molecular mechanism studies have focused on key signalling pathways, such as Wnt/b-catenin signalling. HCC organoid models have demonstrated that axis inhibitor 1 (AXIN1) deficiency drives drug resistance through the WNT/KDM5B/p15 signalling axis. Mechanistically, AXIN1-deficient HCC organoids exhibit enhanced b-catenin nuclear translocation, leading to cellular myelocytomatosis oncogene (c-Myc) overexpression. The subsequent formation of a c-Myc/Miz-1 transcriptional repressor complex mediates p15 epigenetic silencing via KDM5B-dependent H3K4 demethylation (H3K4me2↓). This molecular mechanism results in a > 3-fold reduction in lenvatinib sensitivity in organoids, faithfully recapitulating the drug-resistant phenotype observed in clinical AXIN1-mutant HCC cases [23]. Another similar study suggested that increased expression of the FZD10 gene can promote the expansion of HCC stem cells and lenvatinib resistance by regulating the b-catenin/c-Jun/MEK/ERK axis [24].
Furthermore, the application of HCC organoids in drug development and treatment also occupies an important position, especially in cancer-targeted drug therapy and drug resistance. HCC organoids are primarily used to model clinical challenges, particularly sorafenib resistance and lenvatinib response. Studies reveal that sorafenib resistance involves GSTA1/CTNNB1-mediated ferroptosis suppression through peroxidase activity, while organoid-based screening identifies histone deacetylase inhibitors as potential lenvatinib sensitisers. These findings validate organoids for resistance modelling and highlight new therapeutic targets [25, 26]. These research hotspots were highly consistent with the keywords ‘drug resistance’ and ‘targeted therapy’.
However, studies have shown that the combination of organoids and microfluidic chips can simulate the TME for high-throughput drug screening, providing a new platform for predicting the immunotherapy response of patients with HCC [27]. Another study showed that HCC organoids combined with CRISPR screening technology revealed the potential of combining the clinically approved drugs ifenprodil and sorafenib as a new effective treatment regimen [28]. This multi-technology combination strategy significantly improves research efficiency.
Key word burst analysis showed that ‘long-term expansion’, ‘fibrosis’, ‘patient-derived organoids’, ‘oxidative stress’, ‘primary liver cancer’, ‘b-catenin’, and ‘acid’ have been the fields of continuous concern over the past 2 years. The burst of these keywords strongly indicates future research trends in HCC organoids.
At present, the existing organoid culture still faces problems such as long-term maintenance difficulties and cell dedifferentiation; therefore, the long-term expansion of organoids is particularly important. A research team has created a more physiologically relevant organ model using biomimetic ECM scaffolds to improve the internal microenvironment of organoids [29]. Some studies have shown that by establishing a biobank of organoids, sample accessibility and experimental repeatability can be improved, while also enabling long-term expansion of organoid cultures [30].
Tumour heterogeneity is a key challenge in HCC research. Therefore, it was necessary to refine the research model. Recent studies have found that different gene mutations (such as CTNNB1 mutations or TP53 deletions) can alter tumour characteristics [31, 32]. This suggests that we need to construct a research model with precise genotype-phenotype correspondence to deeply understand the unique biological behaviour of each subtype. Patient-derived organoids are individualised organs that are homologous to the patient’s tumour and are cultured in the laboratory through certain interventional means to obtain the patient’s tumour cells to achieve individualised precision medicine. At present, patient-derived tumour organoids (PDOs) have been used for drug prediction, but further methods to solve individual differences and standardisation problems need to be explored [33].
The keyword ‘b-catenin’ illustrates that the Wnt/b-catenin signalling pathway and targeted therapy for this pathway still face major challenges. It is worth mentioning that the Wnt/b-catenin pathway is activated in 30–40% of HCC, but the targeting strategy is limited. Therefore, new methods for inhibiting this pathway are worthy of further investigation. Encouragingly, a research team has recently made a breakthrough in the development of innovative peptide drugs that effectively block the CCDC137-LZTS2 interaction by specifically targeting the 1–75 domain of the CCDC137 protein, thereby significantly inhibiting the activity of b-catenin and AKT signalling pathways [34].
This study has some limitations that need to be explained. First, the data source is limited to the WOSCC database. Although data consistency is guaranteed, important documents in other databases can be missed. Second, there may be selection bias in the literature screening process, and emerging research directions may not be fully included. In addition, the bibliometric method mainly relies on quantitative indicators, such as the number of publications and citations, and it is difficult to comprehensively evaluate low-cited but high-value research results. It should be emphasised that these limitations do not affect the main conclusions of this study, but it is necessary to maintain a cautious attitude when interpreting the results.
Conclusions
This study reveals the development track of HCC organoids in the past 20 years through the bibliometric analysis system. The results show that China leads with 128 articles (47.94%), but the original articles of European and American institutions in high-impact journals such as Nature and Cell are more prominent. Since 2016, the field has entered a stage of rapid development. The organoids of HCC have gradually developed from simple model construction and optimisation in the early stage to multiple directions such as molecular mechanism exploration, drug screening, targeted therapy, drug resistance mechanism research, and individualised treatment. This is the current research hotspot and research trend in the field. However, its further development is still limited by challenges such as long-term training stability, standardised operating procedures, and insufficient interdisciplinary collaboration. In the future, more in-depth research is needed. In summary, the value of this study is to reveal the evolution law of organoid research of HCC for the first time through bibliometric methods, and provide an objective basis for future research directions.