Introduction
Cancer causes 9.6 million deaths each year, and an estimated 70% of cancer deaths occur in developing countries and is the second leading cause of death in the world [1]. Cancer patients are mostly diagnosed with distant metastases at the time of diagnosis. Currently, cancer metastasis remains an incurable condition with a relatively low survival rate, so novel therapeutic strategies for cancer are needed [2]. Human epidermal growth receptor-2 (HER-2) signalling has become a promising area for the development of molecular targeted therapy for cancer. Several chemotherapeutic agents targeting HER-2 receptors have been developed. Trastuzumab was the first antibody found to target HER-2. However, 70% of patients who use trastuzumab for an extended period develop resistance, leading to the progression of metastatic cancer, primarily due to increased epidermal growth factor receptor (EGFR) expression [3, 4]. The rising incidence of drug resistance in cancer patients poses a significant challenge to both conventional chemotherapy treatment and targeted therapy [5].
Therefore, natural compounds may serve as alternative agents to overcome this resistance. Recent evidence indicates that phytochemicals and natural products play a crucial role in modulating various cancer signalling pathways related to proliferation, apoptosis, and drug resistance. Different groups of bioactive compounds, such as flavonoids, alkaloids, terpenoids, and polyphenols, have been reported to target the PI3K/Akt/mTOR and p53 pathways, as well as regulate reactive oxygen species (ROS) production, thereby contributing to the inhibition of breast cancer cell growth, including the HER-2-positive subtype [6–9]. The integration of natural product-based therapies not only offers opportunities to overcome resistance to conventional treatments but also provides a more comprehensive multimodal approach to cancer management. Several recent studies support this view, emphasizing the potential of natural products in the regulation of cancer signalling pathways [10–12].
Fangchinoline is a compound derived from the dried root of Stephaniae tetrandrine S. Moore, a plant from the Menispermaceae family, traditionally used in Chinese medicine [13–15]. This bisbenzylisoquinoline alkaloid has demonstrated significant anti-cancer properties across various tumour cell lines, including MDA-MB-231 breast cancer cells, MG63 and U20S bone cancer cells, A549 lung cancer cells, PC3 human prostate cancer cells, K562 myelogenous leukaemia cells, T24 and 5637 bladder cancer cells, AGS human gastric cancer cells, and U87 MG and U118 MG glioblastoma multiforme cancer cells [16–20]. Although several studies have reported the anticancer properties of fangchinoline in diverse cancer cell types, limited attention has been given to its role in HER-2-overexpressing breast cancer. To date, no previous studies have combined network pharmacology, molecular docking, molecular dynamics simulations, and in vitro validation to investigate fangchinoline as a potential HER-2 inhibitor. Therefore, this study was designed to provide mechanistic insights into the activity of fangchinoline in HER-2-positive breast cancer.
Specifically, we evaluated the anticancer effects of fangchinoline on MCF-7/HER-2 breast cancer cells, including cytotoxicity, cell cycle regulation, apoptosis, expression of PI3K, Akt, mTOR, p53, and HER-2, as well as its impact on ROS production.
Material and methods
Fangchinoline was purchased from TCI (CAS RN 436-77-1), annexin-V from BioLegend, anti-Akt APC from BioLegend, anti ERBB2/HER-2 from BioLegend, anti-PI3K FITC from Bioss, anti p53 FITC from BioLegend, anti-mTOR PE from BioLegend, distilled water from BrataChem, DMEM from Gibco, DMSO from Sigma, ethanol from Merck, Foetal Bovine Serum from Gibco, propidium iodide from BioLegend, and 3-(4,5-dimethylthiazole-2-yl)-2,5-diphenyl tetrazolium bromide (MTT) from Sigma.
Target identification and annotation
Proteins related to HER-2 positive breast cancer were identified using GeneCards (“human EGFR 2 positive breast cancer”) and DisGeNET (“HER-2 positive breast carcinoma, C1960398”), focusing on protein-coding genes. Fangchinoline targets were predicted via Swiss Target Prediction using isomeric SMILES from PubChem. Intersecting genes were identified using a Venn diagram tool (https://bioinformatics.psb.ugent.be/webtools/Venn/), annotated with DAVID (https://david.ncifcrf.gov/), and visualized using SRplot (https://www.bioinformatics.com.cn/en).
Network construction and analysis
Gene and fangchinoline target lists from SwissTargetPrediction were integrated into a network using Cytoscape 3.10.2. Common targets in HER-2-positive breast cancer were identified and analysed using the CytoHubba plugin based on Betweenness, Closeness, and Degree algorithms. The top three ranked targets were selected for molecular docking and dynamics analysis.
Structure retrieval and molecular preparation
Three-dimensional (3D) protein structures were retrieved from the RCSB protein data bank (PDB), and the structure of fangchinoline (CID: 73481) was obtained from PubChem. Native ligands were extracted from ADRB2 (PDB ID: 3NYA), ERBB2 (3RCD), and IGF1R (2OJ9) using PyMOL, selected based on biological relevance. Protein structures were prepared by removing water molecules and adding missing hydrogen atoms using Avogadro. Geometry optimization and energy minimization were performed with XTB (extreme – opt level), and .xyz files were converted to .pdb using PyMOL.
Molecular docking
Molecular docking and binding affinity calculations were performed using Vina 1.2.5 [21] via the PyRx interface [22], with ligands treated as flexible and proteins as rigid [23]. Docking targeted each protein’s native ligand binding site using grid parameters is shown in Table 1 and an exhaustiveness level of 24. The best conformation was selected based on the lowest root mean square eviation (RMSD). Ligand- protein interactions were visualized with PyMOL and analysed using Biovia Discovery Studio 2019. The top-ranked complex was used for molecular dynamics simulation.
Molecular dynamics
The selected complex was subjected to a 50 ns molecular dynamics simulation using YASARA 23.9.29.W.64 under physiological conditions (pH 7.4, 0.9% NaCl, 310 K, 1 bar) with the AMBER14 force field. Snapshots were recorded every 0.05 ns (total 1000). Post-simulation, RMSD and root mean square fluctuation were calculated to assess structural stability. Free binding energy was estimated using the Poisson-Boltzmann method via the md_analyzebindenergy macro, with surface tension set at 0.65 kJ/mol/Å2 [24, 25].
Cytotoxicity activity
MCF-7/HER-2 cells were cultured in DMEM at a density of 3 × 103 cells/ml and seeded in 0.1 ml portions. After 24 h, the medium was replaced with test samples and incuba- ted for 48 h. Cells were then treated with MTT for 4 h at 37°C, followed by addition of 10% sodium dodecyl sulfate in 0.01 N HCl to stop the reaction. After 24 h incubation in the dark, absorbance was measured at 595 nm and used to calculate cell viability [26].
Cell cycle analysis
MCF-7/HER-2 cells (3 × 105 cells/well) were seeded in 6-well plates and treated with various sample concentrations for 48 h. Cells were fixed with 70% ethanol, stained with propidium iodide/RNase, and analysed for DNA content using a FACScan flow cytometer [9].
Apoptosis analysis
MCF-7/HER-2 cells were cultured in 6-well plates at 3 × 105 cells per well. The cells were then treated with samples for 48 h, collected, cleaned, resuspended in phosphate buffered saline (PBS) solution, and labelled with Annexin V-FITC/PI. The samples were then examined after incubation using a FACScan flow cytometer [27].
PI3K, TOR, p53, Akt, and MCF-7/HER-2 analysis
MCF-7/HER-2 cells (3 × 105 cells/well) were seeded in 6-well plates, incubated for 24 h, and treated for another 48 h. Both adherent and suspended cells were collected using 0.025% trypsin, washed with cold PBS, and centrifuged at 2500 rpm for 5 min. The pellet was incubated with PermWash buffer for 15 min, stained with PI3K FITC, Akt APC, mTOR PE, p53 FITC, and HER-2 FITC antibodies at 37°C for 10 min, and analysed by a FACScan flow cytometer [28].
Reactive oxygen species production analysis
MCF-7/HER-2 cells (3 × 105 cells/well) were seeded in 6-well plates and cultured overnight in DMEM. After harvesting with 0.25% trypsin-EDTA, cells were resuspended in 500 µl 1 × supplement buffer (PBS + 10% FBS), labelled with 25 µM DCFH-DA, and incubated at 37°C with 5% CO2 for 30 min. Samples were then added, followed by a 4 h incubation. Reactive oxygen species levels were measured using a FACScan flow cytometer [29].
Results
Identification of fangchinoline potential targets
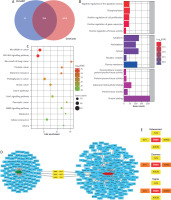
A total of 288 and 9332 genes were retrieved from Gene Cards and DisGeNET, respectively, with 256 overlapping genes (Figure 1 A) potentially linked to HER-2 positive breast cancer. Functional annotation revealed their roles in promoting cell proliferation and inhibiting apoptosis, with predominant expression in cytoplasmic and membrane regions. Most genes were involved in protein binding (Figure 1 B) and associated with multiple cancer pathways (Figure 1 C). Network analysis identified six overlapping targets with fangchinoline: ERBB2, IGF1R, ADRB2, ABCB1, FGFR1, and FLT4 (Figure 1 D). Topology scoring highlighted ERBB2, IGF1R, and ADRB2 as the most significant, which were selected for further analysis.
Figure 1
Identification of fangchinoline target using GeneCards, DisGeNET, and SwissTargetPrediction accompanied by functional annotation and network ranking analysis. A) Venn diagram on the identified targets from GeneCards and DisGeNET. B) Functional annotation on the 256 that co-founded in the GeneCards and DisGeNET databases. C) KEGG pathway classification on the 256 that co-founded in the Gene Cards and DisGeNET databases. D) Network analysis to find the intersection of targets involved in human epidermal growth receptor-2 positive breast cancer and fangchinoline potential targets. E) Ranking algorithm to find the most significant target from the constructed network
FDR – false discovery rate
Interaction of fangchinoline with selected targets
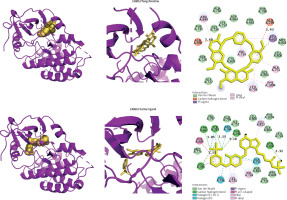
Molecular docking and dynamics were conducted to evaluate fangchinoline’s interaction with selected targets, using native inhibitors as controls. ERBB2 emerged as the most favourable target, with fangchinoline showing a binding energy comparable to the native ligand. In contrast, fangchinoline showed lower affinity for IGF1R and ADRB2 (Table 1, Figure 2), indicating its potential as a selective ERBB2 inhibitor.
Figure 2
The three-dimensional visualization of the ERBB2/fangchinoline and ERBB2/native ligand complexes, along with two-dimensional schematic diagrams illustrating the amino acid-ligand interactions for both complexes, is provided. The distances of hydrogen bonds are indicated in angstroms
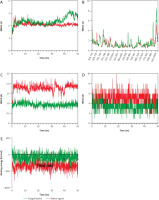
Molecular dynamics simulations confirmed the stability of both ERBB2/fangchinoline and ERBB2/native ligand complexes (Figure 3 A). Residue fluctuations were minimal and did not affect the binding site (Figure 3 B). Fangchinoline exhibited a more stable binding conformation than the native ligand (Figure 3 C), suggesting higher binding affinity. Although it formed fewer hydrogen bonds (Figure 3 D), its binding energy remained comparable, supporting docking results that indicate strong interaction with ERBB2 (Figure 3 E).
Figure 3
Molecular simulations depicting the stability of Erb2/fangchinoline against ERBB2/Native Ligand under physiological conditions. A) Stability of the complexes according to the root mean square deviation of backbone atoms. B) Stability of the protein structures based on the root mean square fluctuation values. C) Ligand conformations’ stability during its interaction with ERBB2. D) Number of formed hydrogen bonds between ligands and ERBB2. E) Free binding energy estimations of ERBB2/fangchinoline compared to ERBB2/Native ligand
RMSD – root mean square deviation
Cytotoxicity activity
In the treatment with fangchinoline and doxorubicin, the values obtained were 9.67 ±0.14 µM and 0.14 ±0.002 µM, respectively. Fangchinoline is classified as an active compound and is considered suitable for targeting malignant cells, as its IC50 value is ≤ 10 µM.
Analysis of cell cycle inhibition
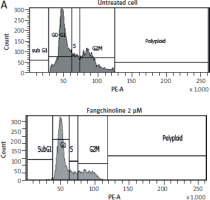
Cell cycle analysis (Figure 4) showed that fangchinoline at 2 and 5 µM induced cell accumulation in the G2-M phase compared to untreated controls, indicating G2-M arrest as the primary inhibitory effect.
Apoptosis analysis
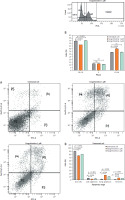
The results of the apoptosis-inducing test on MCF-7/ HER-2 cells are presented in Figure 5. The induction of apoptosis is analysed to determine the causes of cell death, including both apoptosis and necrosis. The flow cytometry method can differentiate live cells, early apoptosis, late apoptosis, and necrosis by staining with Annexin V and Propidium Iodide. The mechanism of cell death can be assessed based on the total percentage in each quadrant. The quadrants are divided into four: quadrant I (lower left) indicates the number of living cells; quadrant II (lower right) indicates early apoptosis; quadrant III (top right) shows cells that have undergone apoptosis and quadrant IV (top left) indicates the number of cells that have undergone necrosis. The results demonstrate that fangchinoline increased early apoptosis, late apoptosis/early necrosis and even late necrosis in MCF-7/HER-2 cells.
Analysis of p53, Akt, PI3K, and mTOR expressions
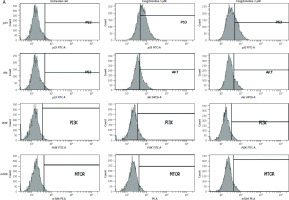
The ability of fangchinoline to inhibit the expression of PI3K, Akt, and mTOR while increasing p53 expression is shown in Figure 6.
Reactive oxygen species production analysis
The ability of fangchinoline to inhibit ROS production is shown in Figure 7.
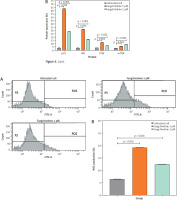
Figure 7
Analysis of reactive oxygen species (ROS) production in MCF-7/HER-2 cells by flow cytometry (A) and percentage of ROS production (B)
ROS – reactive oxygen species Data are presented as mean ±SD (n = 3). p < 0.001 indicates statistical significance and not statistically significant indicates not significant
Human epidermal growth receptor-2 expression analysis
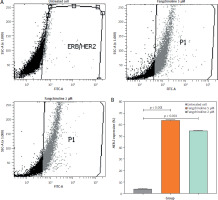
Human epidermal growth receptor-2 expression in cells treated with 2 and 5 µM fangchinoline increased significantly (54.9 and 64.0%, respectively) compared to control cells, as shown in Figure 8.
Discussion
This study utilized network analysis to identify fangchinoline targets relevant to breast cancer, focusing on biological processes such as cell proliferation, signal transduction, and cell death. From GeneCards and DisGeNET, six overlapping targets were identified: ERBB2, IGF1R, ABCB1, FGFR1, ADRB2, and FLT4. Among them, ERBB2, IGF1R, and ADRB2 emerged as the most promising based on network topology analysis. ERBB2 (HER-2), frequently overexpressed in breast cancer, is linked to invasion, metastasis, and chemoresistance [30–32], and is a key therapeutic target to improve disease-free survival [33, 34]. ADRB2 plays a role in cancer progression, with knockdown reducing proliferation and migration [35], while also modulating the tumour microenvironment [36–38]. IGF1R’s prognostic role remains debated, though it holds potential for targeting cell proliferation, despite concerns regarding glucose metabolism disruption [39–41].
Fangchinoline showed lower binding affinity for IGF1R and ADRB2 compared to native ligands but exhibited comparable affinity for ERBB2, binding at similar sites (Table 1, Figure 2). Molecular dynamics analysis supported its stable interaction with ERBB2, indicating potential inhibitory activity (Figure 3). This suggests that fangchinoline may exert antiproliferative effects by targeting ERBB2. ERBB2 has long been a key therapeutic target in breast cancer, with treatments including monoclonal antibodies (e.g., trastuzumab, pertuzumab) and small-molecule inhibitors (e.g., lapatinib, neratinib) [42–45]. Fangchinoline has the potential to suppress ERBB2 activity, making it a promising candidate for anti-breast cancer treatment, particularly in HER-2-positive cases.
Fangchinoline induced G2-M cell cycle arrest in MCF-7/ HER-2 cancer cells (Figure 4), consistent with reports stating that bisbenzylisoquinoline alkaloid compounds increase the accumulation of cells in the G2-M phase of the cell cycle [46, 47]. The key mechanism contributing to the anticancer effect is stopping the progression of the cell cycle by targeting specific proteins, which causes the accumulation of cancer cells at a specific phase. Cell cycle arrest prevents cancer cells from developing into tumours and stops them from spreading [48].
Fangchinoline increased early apoptosis, late apoptosis/ early necrosis and even late necrosis in MCF-7/HER-2 cells (Figure 5). Apoptosis and necrosis are important for understanding cancer development and its treatment. Apoptosis is planned cell death, while necrosis is unintentional cell death. Apoptosis occurs through genetic mechanisms, such as DNA damage or fragmentation. Apoptosis is carried out by caspases, which cleave target proteins. Inhibition of apoptosis can cause cancer. Cells that undergo necrosis due to combination treatment may be affected by accelerated evidence of cell apoptosis early and progress to necrosis [49].
Examination using FACScan flow cytometry showed increased expression of PI3K, mTOR, and Akt in control cells (Figure 6). This indicates that their expression is generally inhibited or suppressed by fangchinoline. The aberrant activation of PI3K, mTOR, and Akt pathways is essential, as this pathway plays a vital role in carcinogenesis and tumour cell proliferation [50]. Additionally, p53 expression was found to increase. p53 functions as a tumour suppressor by activa- ting apoptosis in response to DNA damage [51].
Fangchinoline increased ROS production in MCF-7/HER-2 cells (Figure 7). Reactive oxygen species can induce cell senescence and activate several antiapoptotic proteins. Reactive oxygen species levels are also crucial for maintaining growth arrest during the formation of senescent cells. Upon formation, cancer cell genomes produce polyploidy, the ultimate form of senescence. However, higher ROS le-vels exceeding the threshold will lead to cell death [52].
Furthermore, HER-2 expression in cells treated with 2 and 5 µM fangchinoline increased significantly compared to control cells (Figure 8). The high percentage of expression indicates HER-2 inhibition of cells. Human epidermal growth receptor-2 inhibition in therapy is done by blocking the HER-2 receptor in cancer cells. This inhibition can cause decreased cancer cell proliferation and induce cell death. Drugs such as trastuzumab, pertuzumab, fam-trastuzumab deruxtecan-nxki, lapatinib are commonly used to inhibit HER-2 in the treatment of breast cancer and colorectal cancer by inhibiting the cleavage of the basal and active HER-2 ectodomain in cancer cells, reducing cancer cell proliferation, inducing cell death, and suppressing apoptosis [53, 54].
Conclusions
The analysis of potential targets for fangchinoline in breast cancer treatment involves key biological processes, including cell proliferation, signal transduction, and cell death. ERBB2, ADRB2, and IGF1R were identified as potential targets, with molecular dynamics analysis supporting the inhibitory activity of fangchinoline, particularly on ERBB2. Functionally, fangchinoline induced G2-M phase arrest, triggered apoptosis, and reduced the expression of PI3K, Akt, and mTOR proteins while increasing p53 expression, modulating HER-2, and elevating ROS production. Taken together, these findings indicate that fangchinoline is a promising candidate for further development as an anticancer agent in HER-2-positive breast cancer.